CoNet is very memory intensive.Įxport CLASSPATH=$/CoNet.jarĪlias conet="java -Xmx800m be.ac.CooccurrenceAnalyser" You can allocate more memory to java by setting the -Xmx to a higher number according to how much memory you have on your system, ex. Set environment variables (and an alias if you like) in your. I'm assuming you already have miniconda3 or anaconda on your system.Ĭonda create -name cytoscape -c bioconda cytoscape Set up a new conda environment called cytoscape. CoNet3 needs Cytoscape 3 that needs at least Java 7 (openjdk 1.7.0+). I'm assuming you're working in linux at the command line. Step-by-step method for using CoNet with CytoscapeĬheck what version of java you have installed. Here are the steps I took to get everything working right. Since I was having trouble customizing the analysis in R I switched to Cystoscape instead for more control. Using CoNet in R was a little challenging as the documentation is very abbreviated. CytoNet is available as a 'barebonesCoNet' function in the seqgroup package in R, but the full functionality is only available through the CoNet plug-in that works with Cytoscape (Faust and Raes, 2016 Faust, 2021). False positives due to compositional effects are controlled through permutation and bootstrapping. Show Package Contents, which will display the Contents subdirectoryįor example, if you want Cytoscape to initially allocate 2GB of memoryĪnd use up to a maximum of 4GB, edit the Cytoscape.CytoNet is an ensemble network analysis method that combines several similarity and dissimilarity methods into a single tool (Faust et al., 2012). 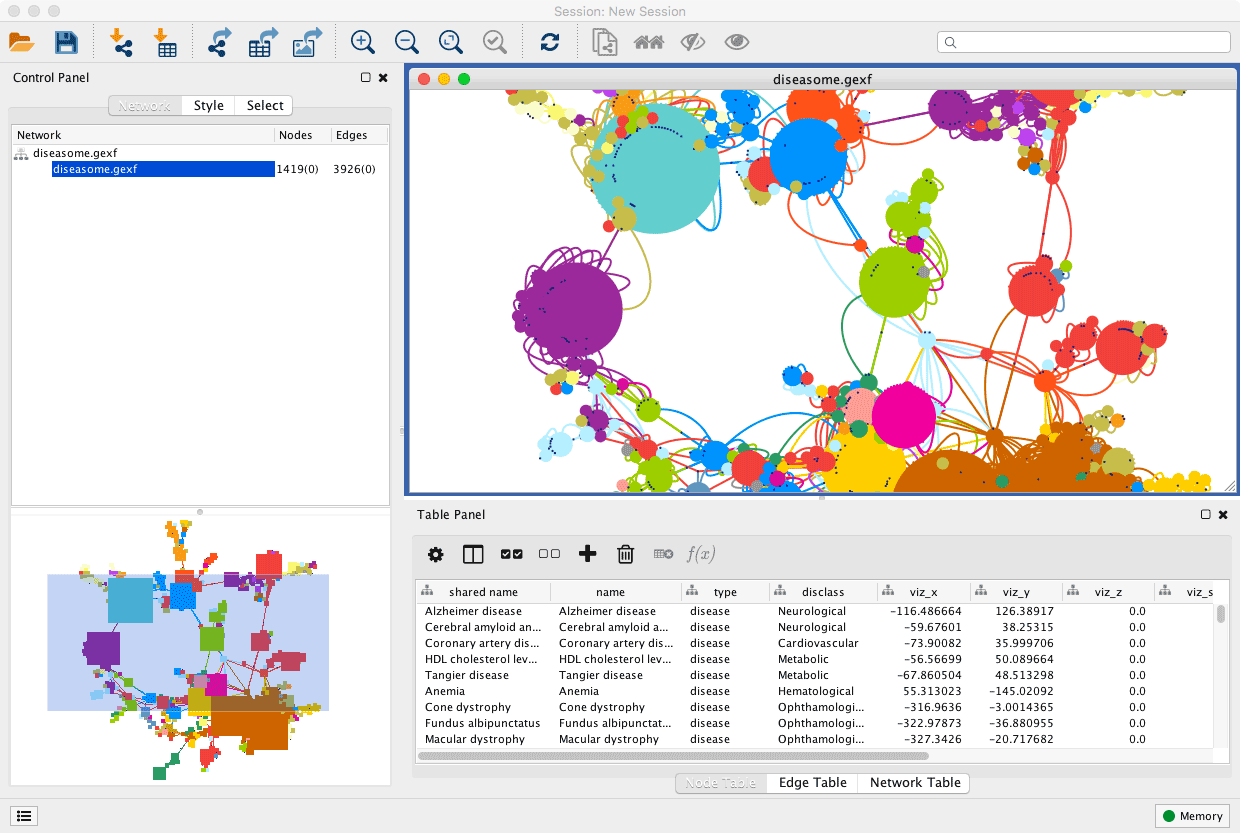
To access this inįinder, you will need to right-click the Cytoscape app icon and select …/Cytoscape.app/Contents/vmoptions.txt file instead. Platform, the situation is slightly different – if you are launchingĬytoscape by clicking on the Cytoscape icon, you must edit the The fileĬontains one option per line, with each line terminated by a linefeed,Īnd an extra linefeed at the end of the file. Resides in the same directory as the Cytoscape executable. Maximum memory size by editing the Cytoscape.vmoptions file, which You can change Cytoscape’s initial and/or Variables – see your operating system documentation for furtherīy default, Cytoscape uses an estimate for initial and maximum memoryĪllocation based on your operating system, system architecture (32 or 64īit), and installed memory. Your operating system may have other mechanisms for setting environment Option to the Cytoscape.vmoptions file or the _JAVA_OPTIONSĮnvironment variable, substituting the desired path as appropriate: Installed on a workstation, but the home directory is stored on aĬentral file server. to the desired directory – this is useful when Cytoscape is To change the user homeĭirectory from the default, one can set the Java environment variable The u/ directory signifies the user’s home directory, which variesįrom user to user and from platform to platform. Various libraries needed to run the application. 
The core CytoscapeĪpplication assumes this directory structure when looking for the For Cytoscape to work properly, all files shouldīe left in the directory in which they were unpacked. The p/ directory signifies the program directory, which varies from U/CytoscapeConfiguration/cytoscape3.props Preset networks as described in the embedded README.txt fileĬytoscape properties and program cache files More automation flexibility is available using other settings and pre-programmed response files, as described in Appendix A of the Install4j manual ().Ĭytoscape installations (regardless of platform) contain theįollowing files and directories: Cytoscape files and directories Directory / FileĬytoscape program files, startup scripts, and default location for session files With a “-q” parameter, the installation package will automatically choose all default settings. For this to succeed, your execution environment must already have sufficient privileges to install software (e.g., for Windows: administrator priveleges). The installation process can be automated and made silent by executing the installation package with the “-q” command parameter (e.g., “Cytoscape_3_8_0-RC1_windows_64bit.exe -q”) from a command line or script. This will bring up a wizard that will lead you through the process, presenting choices for the installation directory, license agreement, file associations and privacy settings. The easiest and most common way to install Cytoscape is by executing an automatic installation package downloaded from the Cytoscape web site.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
